-
PDF
- Split View
-
Views
-
Cite
Cite
Keiki Okazaki, Norikuni Oka, Takuro Shinano, Mitsuru Osaki, Masako Takebe, Differences in the Metabolite Profiles of Spinach (Spinacia oleracea L.) Leaf in Different Concentrations of Nitrate in the Culture Solution, Plant and Cell Physiology, Volume 49, Issue 2, February 2008, Pages 170–177, https://doi.org/10.1093/pcp/pcm173
Close - Share Icon Share
Abstract
The nitrogen (N) status of a plant determines the composition of its major components (amino acids, proteins, carbohydrates and organic acids) and, directly or indirectly, affects the quality of agricultural products in terms of their calorific value and taste. Although these effects are guided by changes in metabolic pathways, no overall metabolic analysis has previously been conducted to demonstrate such effects. Here, metabolite profiling using gas chromatography–mass spectrometry (GC-MS) was used to evaluate the effect of N levels on spinach tissue, comparing two cultivars that differed in their ability to use N. Wide variation in N content was observed without any distinct inhibition of growth in either cultivar. Principal component analysis (PCA) and self-organizing mapping (SOM) were undertaken to describe changes in the metabolites of mature spinach leaves. In PCA, the first component accounted for 44.5% of the total variance, the scores of which was positively correlated with the plant's N content, and a close relationship between metabolite profiles and N status was observed. Both PCA and SOM revealed that metabolites could be broadly divided into two types, correlating either positively or negatively with plant N content. The simple and co-coordinated metabolic stream, containing both general and spinach-specific aspects of plant N content, will be useful in future research on such topics as the detection of environmental effects on spinach through comprehensive metabolic profiling.
Introduction
Nitrogen (N) supply is one of the major environmental factors that regulate plant components and is closely related to crop quality. In spinach (Spinacia oleracea L.), effects of N supply on the content of nitrate, oxalic acid, carbohydrate, ascorbic acid and other antioxidants have been reported (Libert and Franceschi 1987, Elia et al. 1998, Zornoza and Gonzalez 1998, Logan et al. 1999, Santamaria et al. 1999, Ter Steege et al. 1999). Moreover, nitrate is known to act as a signal regulating the activity of many enzymes and transporters, including nitrate reductase, phosphoenolpyruvate carboxylase, malate dehydrogenase, sucrose phosphate synthase and the nitrate transporter involved in regulating carbon (C) and N balance in plants (Scheible et al. 1997). Furthermore, transcriptome analyses have identified mRNAs regulated by nitrate, including those encoding key enzymes in electron transport and pathways involved in the synthesis of trehalose, lipids, alkaloids and phenyl propanoids (Wang et al. 2003, Scheible et al. 2004). Thus, nitrate levels in a nutrient solution may affect not only the pathways that lead to synthesis of specific compounds (e.g. amino acids, amides and organic acids) but also other pathways including secondary metabolites.
Therefore, it is likely that the overall metabolic adaptation of a plant is affected by the N status, and in turn the C/N status. Although there have been many studies into the effect of N application on the level of specific compounds such as free amino acids/amides and nitrate (Haynes and Goh 1978, Darral and Wareing 1981, Barneix and Causin 1996), the effect of N status on a plant's overall metabolism has not been investigated so far.
To detect changes in overall metabolism, a comprehensive and detailed analysis is required. Metabolite profiling using gas chromatography–mass spectrometry (GC-MS) was first successfully applied to plant biology by Roessner et al. (2000). With a freely available database for the identification of metabolites (Kopka et al. 2005), GC-MS-based metabolite profiling has already proven to be a convenient and powerful tool, for example in measuring broad-scale metabolites to distinguish between silent plant phenotypes (Weckwerth et al. 2004), characterizing the response to nutrient deprivation (Bölling and Fiehn 2005, Nikiforova et al. 2005) and investigating regulatory aspects of fruit development (Carrari et al. 2006). The method has extreme sensitivity in detecting both genetic and environmental effects on biological systems (Fiehn 2002). Moreover, its application is not restricted to model plants, but can be used with any species, including those of agricultural and economic importance.
In this study, we used GC-MS-based metabolite profiling in mature spinach leaves to measure a total of 51 metabolites, comprising sugars, organic acids and amino acids (designated as primary metabolites). The effect of N level—manipulated by changing the concentration of nitrate in the culture solution—on these metabolites was investigated, characterizing the change of primary metabolites on a broad scale. Two cultivars with different capacities for N utilization were chosen, in order to investigate the effect of N on broader nutritional levels and to identify common metabolites responding to N in two cultivars.
Results
Plant growth
Twenty-six-day-old seedlings of cultivars Sanpia or Spade one were treated with varying concentrations of N in the culture medium for a further 8 d prior to harvest. N concentrations ranged from 4 mmol NO3 (denoted 4N) down to 1 mmol NO3 (1N). At harvest, fresh weight fell with decreasing N supply in both cultivars (Table 1); this decrease was statistically significant in the case of cv. Sanpia. While both fresh weight and total N content fell with decreasing N supply, the appearance of the plants did not indicate any stress resulting from N deficiency. As we wanted to investigate the continuous change in metabolites under stress-free conditions, the samples obtained were therefore considered suitable for further experimentation. The decrease in plant growth and total N content was more noticeable in cv. Sanpia, reflecting its greater ability to use N.
Fresh weight (FW; g plant−1), dry matter (DM) ratio (dry weight/fresh weight), total nitrogen concentration (g FW−1), and NO3-N concentration (g FW−1) in mature leaves of spinach, cv. Sanpia and cv. Spade one, grown at different nitrogen concentrations
| Exp . | g FW . | DM ratio . | Total N mg kg−1 FW . | NO3-N mg kg−1 FW . | |
|---|---|---|---|---|---|
| Sanpia | |||||
| 1N | 30.7 ± 1.6b | 0.109 ± 0.002a | 30.3 ± 0.7a | 0.1 ± 0.0a | |
| 2N | 44.3 ± 1.7c | 0.093 ± 0.002b,c | 41.3 ± 0.6b | 2.4 ± 0.3b | |
| 4N | 56.5 ± 4.1d | 0.087 ± 0.003c | 49.7 ± 0.6c | 11.0 ± 0.8d | |
| Spade one | |||||
| 1N | 22.0 ± 0.5a | 0.098 ± 0.001b | 41.6 ± 1.0b | 1.1 ± 0.2a,b | |
| 2N | 25.5 ± 0.9a,b | 0.093 ± 0.001b,c | 48.5 ± 0.7c | 6.8 ± 0.4c | |
| 4N | 25.6 ± 0.7a,b | 0.093 ± 0.002b,c | 49.7 ± 0.6c | 10.5 ± 0.7d |
| Exp . | g FW . | DM ratio . | Total N mg kg−1 FW . | NO3-N mg kg−1 FW . | |
|---|---|---|---|---|---|
| Sanpia | |||||
| 1N | 30.7 ± 1.6b | 0.109 ± 0.002a | 30.3 ± 0.7a | 0.1 ± 0.0a | |
| 2N | 44.3 ± 1.7c | 0.093 ± 0.002b,c | 41.3 ± 0.6b | 2.4 ± 0.3b | |
| 4N | 56.5 ± 4.1d | 0.087 ± 0.003c | 49.7 ± 0.6c | 11.0 ± 0.8d | |
| Spade one | |||||
| 1N | 22.0 ± 0.5a | 0.098 ± 0.001b | 41.6 ± 1.0b | 1.1 ± 0.2a,b | |
| 2N | 25.5 ± 0.9a,b | 0.093 ± 0.001b,c | 48.5 ± 0.7c | 6.8 ± 0.4c | |
| 4N | 25.6 ± 0.7a,b | 0.093 ± 0.002b,c | 49.7 ± 0.6c | 10.5 ± 0.7d |
1N, 2N and 4N denote 1, 2 and 4 mmol NO3 in the culture solution, respectively. Means ± SE (n = 6). Values within one column (Sanpia and Spade one) followed by different letters were significantly different by Tukey's t-test (P < 0.05).
Fresh weight (FW; g plant−1), dry matter (DM) ratio (dry weight/fresh weight), total nitrogen concentration (g FW−1), and NO3-N concentration (g FW−1) in mature leaves of spinach, cv. Sanpia and cv. Spade one, grown at different nitrogen concentrations
| Exp . | g FW . | DM ratio . | Total N mg kg−1 FW . | NO3-N mg kg−1 FW . | |
|---|---|---|---|---|---|
| Sanpia | |||||
| 1N | 30.7 ± 1.6b | 0.109 ± 0.002a | 30.3 ± 0.7a | 0.1 ± 0.0a | |
| 2N | 44.3 ± 1.7c | 0.093 ± 0.002b,c | 41.3 ± 0.6b | 2.4 ± 0.3b | |
| 4N | 56.5 ± 4.1d | 0.087 ± 0.003c | 49.7 ± 0.6c | 11.0 ± 0.8d | |
| Spade one | |||||
| 1N | 22.0 ± 0.5a | 0.098 ± 0.001b | 41.6 ± 1.0b | 1.1 ± 0.2a,b | |
| 2N | 25.5 ± 0.9a,b | 0.093 ± 0.001b,c | 48.5 ± 0.7c | 6.8 ± 0.4c | |
| 4N | 25.6 ± 0.7a,b | 0.093 ± 0.002b,c | 49.7 ± 0.6c | 10.5 ± 0.7d |
| Exp . | g FW . | DM ratio . | Total N mg kg−1 FW . | NO3-N mg kg−1 FW . | |
|---|---|---|---|---|---|
| Sanpia | |||||
| 1N | 30.7 ± 1.6b | 0.109 ± 0.002a | 30.3 ± 0.7a | 0.1 ± 0.0a | |
| 2N | 44.3 ± 1.7c | 0.093 ± 0.002b,c | 41.3 ± 0.6b | 2.4 ± 0.3b | |
| 4N | 56.5 ± 4.1d | 0.087 ± 0.003c | 49.7 ± 0.6c | 11.0 ± 0.8d | |
| Spade one | |||||
| 1N | 22.0 ± 0.5a | 0.098 ± 0.001b | 41.6 ± 1.0b | 1.1 ± 0.2a,b | |
| 2N | 25.5 ± 0.9a,b | 0.093 ± 0.001b,c | 48.5 ± 0.7c | 6.8 ± 0.4c | |
| 4N | 25.6 ± 0.7a,b | 0.093 ± 0.002b,c | 49.7 ± 0.6c | 10.5 ± 0.7d |
1N, 2N and 4N denote 1, 2 and 4 mmol NO3 in the culture solution, respectively. Means ± SE (n = 6). Values within one column (Sanpia and Spade one) followed by different letters were significantly different by Tukey's t-test (P < 0.05).
Profiling of the identified metabolites
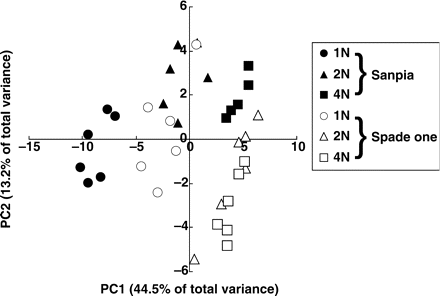
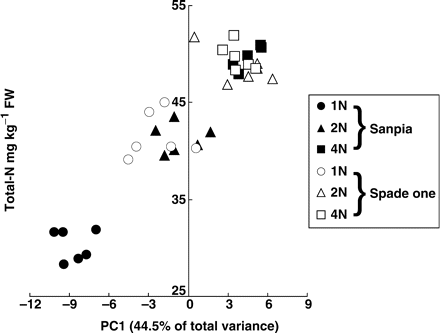
Fifty-one compounds were identified in one scan of GC-MS. Each peak area was subjected to multivariate analysis. Plots of the first and second principal component analysis (PCA) scores revealed the differences in metabolic profiles: the results formed distinct clusters that clearly corresponded to the differences in N levels and cultivars (Fig. 1). The first factor accounted for 44.5% of the total variance, and the N treatment appears to be separated by this factor. This observation was confirmed by plotting the first-factor score against the N content of the whole plant (Fig. 2). The significant positive relationship seen in Fig. 2 indicates a close interaction between the metabolite profile and plant N content.
Sample scores for the first (PC1) and second (PC2) principal components provided by PCA analysis for identified metabolites in spinach leaf extracts. Each group is represented by 5–6 samples. r = 0.921.
Levels of total nitrogen in spinach leaf with PC1 scores, growing at 1, 2 and 4 mM nitrate. Each group is represented by 5–6 samples. r = 0.921.
By using loading data from the first factor, each identified compound was assigned to one of two categories (Table 2). In tissues richer in N, compounds with a negative loading score, such as glucose, fructose, sucrose, lysine and tryptophan, tended to be present in lower concentrations. In contrast, compounds with a positive loading score, such as most amino acids, organic acids and cinnamic acids, tended to be present in higher concentrations.
Levels, self-organizing mapping (SOM) cluster numbers and principal component analysis (PCA) loadings of metabolites with first principal component (PC1) and second principal component (PC2) in spinach leaves, cv. Sanpia and cv. Spade one, grown at different nitrate concentrations
| . | PCA loadings . | SOM cluster member- ship . | Sanpia . | Spade one . | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| . | PC1 . | PC2 . | . | 1N . | 2N . | 4N . | P-value . | 1N . | 2N . | 4N . | P-value . |
| l-Tryptophan | −0.158 | 0.137 | C2 | 1.00 ± 0.24 | 0.78 ± 0.28 | 0.38 ± 0.05 | 0.001* | 0.89 ± 0.19 | 0.42 ± 0.08 | 0.40 ± 0.04 | 0.000* |
| Sucrose | −0.140 | 0.150 | C2 | 1.00 ± 0.14 | 0.80 ± 0.07 | 0.47 ± 0.04 | 0.000* | 0.98 ± 0.18 | 0.59 ± 0.12 | 0.53 ± 0.10 | 0.000* |
| Glucose | −0.132 | 0.201 | C2 | 1.00 ± 0.31 | 1.24 ± 0.59 | 0.31 ± 0.20 | 0.007* | 0.31 ± 0.08 | 0.15 ± 0.07 | 0.13 ± 0.06 | 0.001* |
| Xylose | −0.131 | 0.225 | C6 | 1.00 ± 0.11 | 0.85 ± 0.17 | 0.74 ± 0.06 | 0.014* | 0.76 ± 0.18 | 0.51 ± 0.12 | 0.42 ± 0.09 | 0.002* |
| Arabinose | −0.118 | 0.270 | C6 | 1.00 ± 0.15 | 1.08 ± 0.16 | 0.79 ± 0.11 | 0.012* | 0.84 ± 0.20 | 0.41 ± 0.10 | 0.42 ± 0.06 | 0.000* |
| Fructose | −0.110 | 0.236 | C2 | 1.00 ± 0.49 | 1.94 ± 0.78 | 0.44 ± 0.25 | 0.002* | 0.50 ± 0.07 | 0.19 ± 0.08 | 0.21 ± 0.07 | 0.000* |
| myo-Inositol | −0.094 | 0.226 | C2 | 1.00 ± 0.13 | 0.77 ± 0.08 | 0.71 ± 0.06 | 0.000* | 0.70 ± 0.18 | 0.71 ± 0.13 | 0.56 ± 0.08 | 0.143 |
| l-Lysine | −0.079 | 0.239 | C2 | 1.00 ± 0.13 | 1.18 ± 0.52 | 0.87 ± 0.22 | 0.349 | 1.03 ± 0.39 | 0.62 ± 0.23 | 0.61 ± 0.14 | 0.027* |
| Erythritol | −0.076 | 0.235 | C6 | 1.00 ± 0.13 | 1.07 ± 0.07 | 1.17 ± 0.14 | 0.074 | 0.17 ± 0.05 | 0.16 ± 0.06 | 0.10 ± 0.02 | 0.055 |
| Ribose | −0.031 | 0.266 | C2 | 1.00 ± 0.21 | 1.24 ± 0.30 | 1.18 ± 0.13 | 0.199 | 1.50 ± 0.35 | 0.93 ± 0.11 | 0.79 ± 0.17 | 0.000* |
| l-Tyrosine | 0.113 | −0.040 | C4 | 1.00 ± 0.51 | 1.97 ± 0.46 | 2.19 ± 0.23 | 0.001* | 1.71 ± 0.15 | 2.60 ± 0.52 | 2.69 ± 0.29 | 0.000* |
| Malate | 0.124 | 0.064 | C1 | 1.00 ± 0.23 | 2.41 ± 1.35 | 3.84 ± 0.55 | 0.000* | 0.75 ± 0.21 | 1.66 ± 0.33 | 2.23 ± 0.61 | 0.000* |
| Benzoate | 0.130 | −0.070 | C5 | 1.00 ± 0.30 | 1.71 ± 0.53 | 2.68 ± 0.95 | 0.002* | 3.76 ± 4.45 | 4.18 ± 2.05 | 3.16 ± 0.80 | 0.826 |
| 2-Oxoglutarate | 0.134 | −0.133 | C5 | 1.00 ± 0.14 | 1.28 ± 0.11 | 1.39 ± 0.19 | 0.001* | 1.47 ± 0.17 | 1.56 ± 0.27 | 1.77 ± 0.32 | 0.151 |
| Glycolate | 0.141 | 0.034 | C5 | 1.00 ± 0.20 | 1.70 ± 0.21 | 1.86 ± 0.63 | 0.004* | 1.77 ± 0.51 | 2.21 ± 0.64 | 1.71 ± 0.37 | 0.221 |
| Monomethyl-P | 0.147 | 0.130 | C3 | 1.00 ± 0.16 | 1.40 ± 0.23 | 1.69 ± 0.22 | 0.000* | 1.58 ± 0.35 | 1.68 ± 0.38 | 1.52 ± 0.34 | 0.755 |
| 4-Hydroxycinnamate | 0.148 | 0.195 | C4 | 1.00 ± 0.27 | 2.09 ± 0.72 | 2.66 ± 0.29 | 0.000* | 1.29 ± 0.47 | 1.76 ± 0.55 | 1.74 ± 0.35 | 0.174 |
| trans-Ferulate | 0.157 | 0.076 | C4 | 1.00 ± 0.55 | 2.47 ± 0.91 | 2.88 ± 0.71 | 0.002* | 2.40 ± 0.91 | 3.16 ± 0.65 | 2.75 ± 0.80 | 0.283 |
| Citrate | 0.170 | −0.073 | C4 | 1.00 ± 0.15 | 1.27 ± 0.34 | 2.21 ± 0.45 | 0.000* | 1.15 ± 0.45 | 2.28 ± 0.40 | 2.56 ± 0.21 | 0.000* |
| l-Isoleucine | 0.173 | 0.000 | C1 | 1.00 ± 0.21 | 1.47 ± 0.24 | 1.58 ± 0.14 | 0.001* | 1.41 ± 0.20 | 1.87 ± 0.36 | 1.87 ± 0.08 | 0.006* |
| trans-p-Coumarate | 0.174 | 0.161 | C4 | 1.00 ± 0.10 | 2.22 ± 0.50 | 2.65 ± 0.40 | 0.000* | 1.62 ± 0.44 | 2.04 ± 0.53 | 2.05 ± 0.25 | 0.160 |
| Erythronate | 0.176 | 0.001 | C1 | 1.00 ± 0.20 | 1.68 ± 0.20 | 2.23 ± 0.36 | 0.000* | 1.69 ± 0.38 | 2.31 ± 0.39 | 2.15 ± 0.47 | 0.050 |
| l-Glycerol-3-P | 0.176 | 0.157 | C4 | 1.00 ± 0.11 | 1.70 ± 0.27 | 2.21 ± 0.27 | 0.000* | 1.50 ± 0.30 | 1.82 ± 0.38 | 1.66 ± 0.26 | 0.235 |
| l-Serine | 0.177 | −0.019 | C1 | 1.00 ± 0.19 | 1.66 ± 0.34 | 1.82 ± 0.31 | 0.001* | 1.45 ± 0.29 | 2.45 ± 0.56 | 2.29 ± 0.17 | 0.001* |
| l-Cysteine | 0.178 | 0.014 | C1 | 1.00 ± 0.29 | 2.03 ± 0.43 | 2.50 ± 0.20 | 0.000* | 1.95 ± 0.42 | 2.62 ± 0.50 | 2.45 ± 0.53 | 0.078 |
| l-Hydroxyproline | 0.180 | 0.041 | C1 | 1.00 ± 0.23 | 1.61 ± 0.46 | 2.55 ± 0.27 | 0.000* | 1.47 ± 0.25 | 2.15 ± 0.47 | 2.15 ± 0.42 | 0.012* |
| l-Glutamine | 0.183 | −0.040 | C4 | 1.00 ± 0.30 | 4.93 ± 1.62 | 27.24 ± 4.49 | 0.000* | 1.40 ± 0.23 | 17.30 ± 5.04 | 17.03 ± 2.18 | 0.000* |
| Fumarate | 0.184 | 0.040 | C1 | 1.00 ± 0.10 | 2.37 ± 0.43 | 4.11 ± 0.24 | 0.000* | 1.42 ± 0.37 | 2.84 ± 0.74 | 3.14 ± 0.36 | 0.000* |
| l-Alanine | 0.184 | −0.011 | C1 | 1.00 ± 0.30 | 2.16 ± 0.55 | 4.40 ± 0.49 | 0.000* | 1.40 ± 0.39 | 3.75 ± 0.95 | 3.97 ± 0.53 | 0.000* |
| Threonate | 0.184 | 0.006 | C1 | 1.00 ± 0.22 | 1.96 ± 0.35 | 2.68 ± 0.32 | 0.000* | 1.89 ± 0.33 | 2.63 ± 0.45 | 2.49 ± 0.49 | 0.021* |
| l-Asparagine | 0.184 | 0.042 | C1 | 1.00 ± 0.34 | 2.55 ± 0.58 | 3.95 ± 0.70 | 0.000* | 1.42 ± 0.25 | 3.27 ± 0.91 | 3.23 ± 0.29 | 0.000* |
| l-Aspartate | 0.185 | −0.041 | C1 | 1.00 ± 0.15 | 2.38 ± 0.28 | 7.54 ± 1.17 | 0.000* | 1.27 ± 0.16 | 6.35 ± 1.60 | 5.67 ± 0.79 | 0.000* |
| Succinate | 0.186 | −0.031 | C1 | 1.00 ± 0.12 | 1.65 ± 0.30 | 2.53 ± 0.34 | 0.000* | 1.44 ± 0.16 | 2.20 ± 0.48 | 2.50 ± 0.33 | 0.000* |
| l-Homoserine | 0.186 | −0.048 | C1 | 1.00 ± 0.24 | 1.99 ± 0.64 | 3.80 ± 0.66 | 0.000* | 1.73 ± 0.62 | 4.40 ± 0.83 | 3.83 ± 0.44 | 0.000* |
| l-Glutamate | 0.187 | 0.114 | C1 | 1.00 ± 0.13 | 1.87 ± 0.24 | 2.37 ± 0.16 | 0.000* | 1.57 ± 0.21 | 2.06 ± 0.43 | 1.88 ± 0.20 | 0.038* |
| l-Threonine | 0.193 | 0.030 | C1 | 1.00 ± 0.17 | 1.83 ± 0.25 | 2.56 ± 0.38 | 0.000* | 1.48 ± 0.23 | 2.49 ± 0.51 | 2.39 ± 0.20 | 0.000* |
| Pyroglutamate | 0.193 | 0.063 | C4 | 1.00 ± 0.09 | 1.98 ± 0.25 | 3.24 ± 0.30 | 0.000* | 1.51 ± 0.31 | 2.56 ± 0.56 | 2.48 ± 0.17 | 0.000* |
| l-Valine | 0.194 | 0.058 | C1 | 1.00 ± 0.19 | 2.04 ± 0.35 | 2.85 ± 0.29 | 0.000* | 1.54 ± 0.24 | 2.66 ± 0.57 | 2.49 ± 0.37 | 0.001* |
| PC1 score | −8.70 ± 0.50 | −0.68 ± 0.64 | 4.55 ± 0.43 | −2.33 ± 0.77 | 4.12 ± 0.87 | 3.80 ± 0.36 | |||||
| PC2 score | −0.39 ± 0.59 | 2.85 ± 0.59 | 1.92 ± 0.43 | 0.40 ± 0.96 | −1.43 ± 0.98 | −3.03 ± 0.62 | |||||
| Sample number | 6 | 6 | 5 | 6 | 6 | 6 | |||||
| . | PCA loadings . | SOM cluster member- ship . | Sanpia . | Spade one . | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| . | PC1 . | PC2 . | . | 1N . | 2N . | 4N . | P-value . | 1N . | 2N . | 4N . | P-value . |
| l-Tryptophan | −0.158 | 0.137 | C2 | 1.00 ± 0.24 | 0.78 ± 0.28 | 0.38 ± 0.05 | 0.001* | 0.89 ± 0.19 | 0.42 ± 0.08 | 0.40 ± 0.04 | 0.000* |
| Sucrose | −0.140 | 0.150 | C2 | 1.00 ± 0.14 | 0.80 ± 0.07 | 0.47 ± 0.04 | 0.000* | 0.98 ± 0.18 | 0.59 ± 0.12 | 0.53 ± 0.10 | 0.000* |
| Glucose | −0.132 | 0.201 | C2 | 1.00 ± 0.31 | 1.24 ± 0.59 | 0.31 ± 0.20 | 0.007* | 0.31 ± 0.08 | 0.15 ± 0.07 | 0.13 ± 0.06 | 0.001* |
| Xylose | −0.131 | 0.225 | C6 | 1.00 ± 0.11 | 0.85 ± 0.17 | 0.74 ± 0.06 | 0.014* | 0.76 ± 0.18 | 0.51 ± 0.12 | 0.42 ± 0.09 | 0.002* |
| Arabinose | −0.118 | 0.270 | C6 | 1.00 ± 0.15 | 1.08 ± 0.16 | 0.79 ± 0.11 | 0.012* | 0.84 ± 0.20 | 0.41 ± 0.10 | 0.42 ± 0.06 | 0.000* |
| Fructose | −0.110 | 0.236 | C2 | 1.00 ± 0.49 | 1.94 ± 0.78 | 0.44 ± 0.25 | 0.002* | 0.50 ± 0.07 | 0.19 ± 0.08 | 0.21 ± 0.07 | 0.000* |
| myo-Inositol | −0.094 | 0.226 | C2 | 1.00 ± 0.13 | 0.77 ± 0.08 | 0.71 ± 0.06 | 0.000* | 0.70 ± 0.18 | 0.71 ± 0.13 | 0.56 ± 0.08 | 0.143 |
| l-Lysine | −0.079 | 0.239 | C2 | 1.00 ± 0.13 | 1.18 ± 0.52 | 0.87 ± 0.22 | 0.349 | 1.03 ± 0.39 | 0.62 ± 0.23 | 0.61 ± 0.14 | 0.027* |
| Erythritol | −0.076 | 0.235 | C6 | 1.00 ± 0.13 | 1.07 ± 0.07 | 1.17 ± 0.14 | 0.074 | 0.17 ± 0.05 | 0.16 ± 0.06 | 0.10 ± 0.02 | 0.055 |
| Ribose | −0.031 | 0.266 | C2 | 1.00 ± 0.21 | 1.24 ± 0.30 | 1.18 ± 0.13 | 0.199 | 1.50 ± 0.35 | 0.93 ± 0.11 | 0.79 ± 0.17 | 0.000* |
| l-Tyrosine | 0.113 | −0.040 | C4 | 1.00 ± 0.51 | 1.97 ± 0.46 | 2.19 ± 0.23 | 0.001* | 1.71 ± 0.15 | 2.60 ± 0.52 | 2.69 ± 0.29 | 0.000* |
| Malate | 0.124 | 0.064 | C1 | 1.00 ± 0.23 | 2.41 ± 1.35 | 3.84 ± 0.55 | 0.000* | 0.75 ± 0.21 | 1.66 ± 0.33 | 2.23 ± 0.61 | 0.000* |
| Benzoate | 0.130 | −0.070 | C5 | 1.00 ± 0.30 | 1.71 ± 0.53 | 2.68 ± 0.95 | 0.002* | 3.76 ± 4.45 | 4.18 ± 2.05 | 3.16 ± 0.80 | 0.826 |
| 2-Oxoglutarate | 0.134 | −0.133 | C5 | 1.00 ± 0.14 | 1.28 ± 0.11 | 1.39 ± 0.19 | 0.001* | 1.47 ± 0.17 | 1.56 ± 0.27 | 1.77 ± 0.32 | 0.151 |
| Glycolate | 0.141 | 0.034 | C5 | 1.00 ± 0.20 | 1.70 ± 0.21 | 1.86 ± 0.63 | 0.004* | 1.77 ± 0.51 | 2.21 ± 0.64 | 1.71 ± 0.37 | 0.221 |
| Monomethyl-P | 0.147 | 0.130 | C3 | 1.00 ± 0.16 | 1.40 ± 0.23 | 1.69 ± 0.22 | 0.000* | 1.58 ± 0.35 | 1.68 ± 0.38 | 1.52 ± 0.34 | 0.755 |
| 4-Hydroxycinnamate | 0.148 | 0.195 | C4 | 1.00 ± 0.27 | 2.09 ± 0.72 | 2.66 ± 0.29 | 0.000* | 1.29 ± 0.47 | 1.76 ± 0.55 | 1.74 ± 0.35 | 0.174 |
| trans-Ferulate | 0.157 | 0.076 | C4 | 1.00 ± 0.55 | 2.47 ± 0.91 | 2.88 ± 0.71 | 0.002* | 2.40 ± 0.91 | 3.16 ± 0.65 | 2.75 ± 0.80 | 0.283 |
| Citrate | 0.170 | −0.073 | C4 | 1.00 ± 0.15 | 1.27 ± 0.34 | 2.21 ± 0.45 | 0.000* | 1.15 ± 0.45 | 2.28 ± 0.40 | 2.56 ± 0.21 | 0.000* |
| l-Isoleucine | 0.173 | 0.000 | C1 | 1.00 ± 0.21 | 1.47 ± 0.24 | 1.58 ± 0.14 | 0.001* | 1.41 ± 0.20 | 1.87 ± 0.36 | 1.87 ± 0.08 | 0.006* |
| trans-p-Coumarate | 0.174 | 0.161 | C4 | 1.00 ± 0.10 | 2.22 ± 0.50 | 2.65 ± 0.40 | 0.000* | 1.62 ± 0.44 | 2.04 ± 0.53 | 2.05 ± 0.25 | 0.160 |
| Erythronate | 0.176 | 0.001 | C1 | 1.00 ± 0.20 | 1.68 ± 0.20 | 2.23 ± 0.36 | 0.000* | 1.69 ± 0.38 | 2.31 ± 0.39 | 2.15 ± 0.47 | 0.050 |
| l-Glycerol-3-P | 0.176 | 0.157 | C4 | 1.00 ± 0.11 | 1.70 ± 0.27 | 2.21 ± 0.27 | 0.000* | 1.50 ± 0.30 | 1.82 ± 0.38 | 1.66 ± 0.26 | 0.235 |
| l-Serine | 0.177 | −0.019 | C1 | 1.00 ± 0.19 | 1.66 ± 0.34 | 1.82 ± 0.31 | 0.001* | 1.45 ± 0.29 | 2.45 ± 0.56 | 2.29 ± 0.17 | 0.001* |
| l-Cysteine | 0.178 | 0.014 | C1 | 1.00 ± 0.29 | 2.03 ± 0.43 | 2.50 ± 0.20 | 0.000* | 1.95 ± 0.42 | 2.62 ± 0.50 | 2.45 ± 0.53 | 0.078 |
| l-Hydroxyproline | 0.180 | 0.041 | C1 | 1.00 ± 0.23 | 1.61 ± 0.46 | 2.55 ± 0.27 | 0.000* | 1.47 ± 0.25 | 2.15 ± 0.47 | 2.15 ± 0.42 | 0.012* |
| l-Glutamine | 0.183 | −0.040 | C4 | 1.00 ± 0.30 | 4.93 ± 1.62 | 27.24 ± 4.49 | 0.000* | 1.40 ± 0.23 | 17.30 ± 5.04 | 17.03 ± 2.18 | 0.000* |
| Fumarate | 0.184 | 0.040 | C1 | 1.00 ± 0.10 | 2.37 ± 0.43 | 4.11 ± 0.24 | 0.000* | 1.42 ± 0.37 | 2.84 ± 0.74 | 3.14 ± 0.36 | 0.000* |
| l-Alanine | 0.184 | −0.011 | C1 | 1.00 ± 0.30 | 2.16 ± 0.55 | 4.40 ± 0.49 | 0.000* | 1.40 ± 0.39 | 3.75 ± 0.95 | 3.97 ± 0.53 | 0.000* |
| Threonate | 0.184 | 0.006 | C1 | 1.00 ± 0.22 | 1.96 ± 0.35 | 2.68 ± 0.32 | 0.000* | 1.89 ± 0.33 | 2.63 ± 0.45 | 2.49 ± 0.49 | 0.021* |
| l-Asparagine | 0.184 | 0.042 | C1 | 1.00 ± 0.34 | 2.55 ± 0.58 | 3.95 ± 0.70 | 0.000* | 1.42 ± 0.25 | 3.27 ± 0.91 | 3.23 ± 0.29 | 0.000* |
| l-Aspartate | 0.185 | −0.041 | C1 | 1.00 ± 0.15 | 2.38 ± 0.28 | 7.54 ± 1.17 | 0.000* | 1.27 ± 0.16 | 6.35 ± 1.60 | 5.67 ± 0.79 | 0.000* |
| Succinate | 0.186 | −0.031 | C1 | 1.00 ± 0.12 | 1.65 ± 0.30 | 2.53 ± 0.34 | 0.000* | 1.44 ± 0.16 | 2.20 ± 0.48 | 2.50 ± 0.33 | 0.000* |
| l-Homoserine | 0.186 | −0.048 | C1 | 1.00 ± 0.24 | 1.99 ± 0.64 | 3.80 ± 0.66 | 0.000* | 1.73 ± 0.62 | 4.40 ± 0.83 | 3.83 ± 0.44 | 0.000* |
| l-Glutamate | 0.187 | 0.114 | C1 | 1.00 ± 0.13 | 1.87 ± 0.24 | 2.37 ± 0.16 | 0.000* | 1.57 ± 0.21 | 2.06 ± 0.43 | 1.88 ± 0.20 | 0.038* |
| l-Threonine | 0.193 | 0.030 | C1 | 1.00 ± 0.17 | 1.83 ± 0.25 | 2.56 ± 0.38 | 0.000* | 1.48 ± 0.23 | 2.49 ± 0.51 | 2.39 ± 0.20 | 0.000* |
| Pyroglutamate | 0.193 | 0.063 | C4 | 1.00 ± 0.09 | 1.98 ± 0.25 | 3.24 ± 0.30 | 0.000* | 1.51 ± 0.31 | 2.56 ± 0.56 | 2.48 ± 0.17 | 0.000* |
| l-Valine | 0.194 | 0.058 | C1 | 1.00 ± 0.19 | 2.04 ± 0.35 | 2.85 ± 0.29 | 0.000* | 1.54 ± 0.24 | 2.66 ± 0.57 | 2.49 ± 0.37 | 0.001* |
| PC1 score | −8.70 ± 0.50 | −0.68 ± 0.64 | 4.55 ± 0.43 | −2.33 ± 0.77 | 4.12 ± 0.87 | 3.80 ± 0.36 | |||||
| PC2 score | −0.39 ± 0.59 | 2.85 ± 0.59 | 1.92 ± 0.43 | 0.40 ± 0.96 | −1.43 ± 0.98 | −3.03 ± 0.62 | |||||
| Sample number | 6 | 6 | 5 | 6 | 6 | 6 | |||||
Selected metabolites represent a high (>0.1) or low (less than −0.01) loading score of PC1. The P-value was determined by ANOVA. *P < 0.05. Compounds in bold indicate significance in both cv. Sanpia and cv. Spade one. In SOM analysis, each compound is assigned to one of the six cluster groups (C1–C6).
Levels, self-organizing mapping (SOM) cluster numbers and principal component analysis (PCA) loadings of metabolites with first principal component (PC1) and second principal component (PC2) in spinach leaves, cv. Sanpia and cv. Spade one, grown at different nitrate concentrations
| . | PCA loadings . | SOM cluster member- ship . | Sanpia . | Spade one . | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| . | PC1 . | PC2 . | . | 1N . | 2N . | 4N . | P-value . | 1N . | 2N . | 4N . | P-value . |
| l-Tryptophan | −0.158 | 0.137 | C2 | 1.00 ± 0.24 | 0.78 ± 0.28 | 0.38 ± 0.05 | 0.001* | 0.89 ± 0.19 | 0.42 ± 0.08 | 0.40 ± 0.04 | 0.000* |
| Sucrose | −0.140 | 0.150 | C2 | 1.00 ± 0.14 | 0.80 ± 0.07 | 0.47 ± 0.04 | 0.000* | 0.98 ± 0.18 | 0.59 ± 0.12 | 0.53 ± 0.10 | 0.000* |
| Glucose | −0.132 | 0.201 | C2 | 1.00 ± 0.31 | 1.24 ± 0.59 | 0.31 ± 0.20 | 0.007* | 0.31 ± 0.08 | 0.15 ± 0.07 | 0.13 ± 0.06 | 0.001* |
| Xylose | −0.131 | 0.225 | C6 | 1.00 ± 0.11 | 0.85 ± 0.17 | 0.74 ± 0.06 | 0.014* | 0.76 ± 0.18 | 0.51 ± 0.12 | 0.42 ± 0.09 | 0.002* |
| Arabinose | −0.118 | 0.270 | C6 | 1.00 ± 0.15 | 1.08 ± 0.16 | 0.79 ± 0.11 | 0.012* | 0.84 ± 0.20 | 0.41 ± 0.10 | 0.42 ± 0.06 | 0.000* |
| Fructose | −0.110 | 0.236 | C2 | 1.00 ± 0.49 | 1.94 ± 0.78 | 0.44 ± 0.25 | 0.002* | 0.50 ± 0.07 | 0.19 ± 0.08 | 0.21 ± 0.07 | 0.000* |
| myo-Inositol | −0.094 | 0.226 | C2 | 1.00 ± 0.13 | 0.77 ± 0.08 | 0.71 ± 0.06 | 0.000* | 0.70 ± 0.18 | 0.71 ± 0.13 | 0.56 ± 0.08 | 0.143 |
| l-Lysine | −0.079 | 0.239 | C2 | 1.00 ± 0.13 | 1.18 ± 0.52 | 0.87 ± 0.22 | 0.349 | 1.03 ± 0.39 | 0.62 ± 0.23 | 0.61 ± 0.14 | 0.027* |
| Erythritol | −0.076 | 0.235 | C6 | 1.00 ± 0.13 | 1.07 ± 0.07 | 1.17 ± 0.14 | 0.074 | 0.17 ± 0.05 | 0.16 ± 0.06 | 0.10 ± 0.02 | 0.055 |
| Ribose | −0.031 | 0.266 | C2 | 1.00 ± 0.21 | 1.24 ± 0.30 | 1.18 ± 0.13 | 0.199 | 1.50 ± 0.35 | 0.93 ± 0.11 | 0.79 ± 0.17 | 0.000* |
| l-Tyrosine | 0.113 | −0.040 | C4 | 1.00 ± 0.51 | 1.97 ± 0.46 | 2.19 ± 0.23 | 0.001* | 1.71 ± 0.15 | 2.60 ± 0.52 | 2.69 ± 0.29 | 0.000* |
| Malate | 0.124 | 0.064 | C1 | 1.00 ± 0.23 | 2.41 ± 1.35 | 3.84 ± 0.55 | 0.000* | 0.75 ± 0.21 | 1.66 ± 0.33 | 2.23 ± 0.61 | 0.000* |
| Benzoate | 0.130 | −0.070 | C5 | 1.00 ± 0.30 | 1.71 ± 0.53 | 2.68 ± 0.95 | 0.002* | 3.76 ± 4.45 | 4.18 ± 2.05 | 3.16 ± 0.80 | 0.826 |
| 2-Oxoglutarate | 0.134 | −0.133 | C5 | 1.00 ± 0.14 | 1.28 ± 0.11 | 1.39 ± 0.19 | 0.001* | 1.47 ± 0.17 | 1.56 ± 0.27 | 1.77 ± 0.32 | 0.151 |
| Glycolate | 0.141 | 0.034 | C5 | 1.00 ± 0.20 | 1.70 ± 0.21 | 1.86 ± 0.63 | 0.004* | 1.77 ± 0.51 | 2.21 ± 0.64 | 1.71 ± 0.37 | 0.221 |
| Monomethyl-P | 0.147 | 0.130 | C3 | 1.00 ± 0.16 | 1.40 ± 0.23 | 1.69 ± 0.22 | 0.000* | 1.58 ± 0.35 | 1.68 ± 0.38 | 1.52 ± 0.34 | 0.755 |
| 4-Hydroxycinnamate | 0.148 | 0.195 | C4 | 1.00 ± 0.27 | 2.09 ± 0.72 | 2.66 ± 0.29 | 0.000* | 1.29 ± 0.47 | 1.76 ± 0.55 | 1.74 ± 0.35 | 0.174 |
| trans-Ferulate | 0.157 | 0.076 | C4 | 1.00 ± 0.55 | 2.47 ± 0.91 | 2.88 ± 0.71 | 0.002* | 2.40 ± 0.91 | 3.16 ± 0.65 | 2.75 ± 0.80 | 0.283 |
| Citrate | 0.170 | −0.073 | C4 | 1.00 ± 0.15 | 1.27 ± 0.34 | 2.21 ± 0.45 | 0.000* | 1.15 ± 0.45 | 2.28 ± 0.40 | 2.56 ± 0.21 | 0.000* |
| l-Isoleucine | 0.173 | 0.000 | C1 | 1.00 ± 0.21 | 1.47 ± 0.24 | 1.58 ± 0.14 | 0.001* | 1.41 ± 0.20 | 1.87 ± 0.36 | 1.87 ± 0.08 | 0.006* |
| trans-p-Coumarate | 0.174 | 0.161 | C4 | 1.00 ± 0.10 | 2.22 ± 0.50 | 2.65 ± 0.40 | 0.000* | 1.62 ± 0.44 | 2.04 ± 0.53 | 2.05 ± 0.25 | 0.160 |
| Erythronate | 0.176 | 0.001 | C1 | 1.00 ± 0.20 | 1.68 ± 0.20 | 2.23 ± 0.36 | 0.000* | 1.69 ± 0.38 | 2.31 ± 0.39 | 2.15 ± 0.47 | 0.050 |
| l-Glycerol-3-P | 0.176 | 0.157 | C4 | 1.00 ± 0.11 | 1.70 ± 0.27 | 2.21 ± 0.27 | 0.000* | 1.50 ± 0.30 | 1.82 ± 0.38 | 1.66 ± 0.26 | 0.235 |
| l-Serine | 0.177 | −0.019 | C1 | 1.00 ± 0.19 | 1.66 ± 0.34 | 1.82 ± 0.31 | 0.001* | 1.45 ± 0.29 | 2.45 ± 0.56 | 2.29 ± 0.17 | 0.001* |
| l-Cysteine | 0.178 | 0.014 | C1 | 1.00 ± 0.29 | 2.03 ± 0.43 | 2.50 ± 0.20 | 0.000* | 1.95 ± 0.42 | 2.62 ± 0.50 | 2.45 ± 0.53 | 0.078 |
| l-Hydroxyproline | 0.180 | 0.041 | C1 | 1.00 ± 0.23 | 1.61 ± 0.46 | 2.55 ± 0.27 | 0.000* | 1.47 ± 0.25 | 2.15 ± 0.47 | 2.15 ± 0.42 | 0.012* |
| l-Glutamine | 0.183 | −0.040 | C4 | 1.00 ± 0.30 | 4.93 ± 1.62 | 27.24 ± 4.49 | 0.000* | 1.40 ± 0.23 | 17.30 ± 5.04 | 17.03 ± 2.18 | 0.000* |
| Fumarate | 0.184 | 0.040 | C1 | 1.00 ± 0.10 | 2.37 ± 0.43 | 4.11 ± 0.24 | 0.000* | 1.42 ± 0.37 | 2.84 ± 0.74 | 3.14 ± 0.36 | 0.000* |
| l-Alanine | 0.184 | −0.011 | C1 | 1.00 ± 0.30 | 2.16 ± 0.55 | 4.40 ± 0.49 | 0.000* | 1.40 ± 0.39 | 3.75 ± 0.95 | 3.97 ± 0.53 | 0.000* |
| Threonate | 0.184 | 0.006 | C1 | 1.00 ± 0.22 | 1.96 ± 0.35 | 2.68 ± 0.32 | 0.000* | 1.89 ± 0.33 | 2.63 ± 0.45 | 2.49 ± 0.49 | 0.021* |
| l-Asparagine | 0.184 | 0.042 | C1 | 1.00 ± 0.34 | 2.55 ± 0.58 | 3.95 ± 0.70 | 0.000* | 1.42 ± 0.25 | 3.27 ± 0.91 | 3.23 ± 0.29 | 0.000* |
| l-Aspartate | 0.185 | −0.041 | C1 | 1.00 ± 0.15 | 2.38 ± 0.28 | 7.54 ± 1.17 | 0.000* | 1.27 ± 0.16 | 6.35 ± 1.60 | 5.67 ± 0.79 | 0.000* |
| Succinate | 0.186 | −0.031 | C1 | 1.00 ± 0.12 | 1.65 ± 0.30 | 2.53 ± 0.34 | 0.000* | 1.44 ± 0.16 | 2.20 ± 0.48 | 2.50 ± 0.33 | 0.000* |
| l-Homoserine | 0.186 | −0.048 | C1 | 1.00 ± 0.24 | 1.99 ± 0.64 | 3.80 ± 0.66 | 0.000* | 1.73 ± 0.62 | 4.40 ± 0.83 | 3.83 ± 0.44 | 0.000* |
| l-Glutamate | 0.187 | 0.114 | C1 | 1.00 ± 0.13 | 1.87 ± 0.24 | 2.37 ± 0.16 | 0.000* | 1.57 ± 0.21 | 2.06 ± 0.43 | 1.88 ± 0.20 | 0.038* |
| l-Threonine | 0.193 | 0.030 | C1 | 1.00 ± 0.17 | 1.83 ± 0.25 | 2.56 ± 0.38 | 0.000* | 1.48 ± 0.23 | 2.49 ± 0.51 | 2.39 ± 0.20 | 0.000* |
| Pyroglutamate | 0.193 | 0.063 | C4 | 1.00 ± 0.09 | 1.98 ± 0.25 | 3.24 ± 0.30 | 0.000* | 1.51 ± 0.31 | 2.56 ± 0.56 | 2.48 ± 0.17 | 0.000* |
| l-Valine | 0.194 | 0.058 | C1 | 1.00 ± 0.19 | 2.04 ± 0.35 | 2.85 ± 0.29 | 0.000* | 1.54 ± 0.24 | 2.66 ± 0.57 | 2.49 ± 0.37 | 0.001* |
| PC1 score | −8.70 ± 0.50 | −0.68 ± 0.64 | 4.55 ± 0.43 | −2.33 ± 0.77 | 4.12 ± 0.87 | 3.80 ± 0.36 | |||||
| PC2 score | −0.39 ± 0.59 | 2.85 ± 0.59 | 1.92 ± 0.43 | 0.40 ± 0.96 | −1.43 ± 0.98 | −3.03 ± 0.62 | |||||
| Sample number | 6 | 6 | 5 | 6 | 6 | 6 | |||||
| . | PCA loadings . | SOM cluster member- ship . | Sanpia . | Spade one . | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| . | PC1 . | PC2 . | . | 1N . | 2N . | 4N . | P-value . | 1N . | 2N . | 4N . | P-value . |
| l-Tryptophan | −0.158 | 0.137 | C2 | 1.00 ± 0.24 | 0.78 ± 0.28 | 0.38 ± 0.05 | 0.001* | 0.89 ± 0.19 | 0.42 ± 0.08 | 0.40 ± 0.04 | 0.000* |
| Sucrose | −0.140 | 0.150 | C2 | 1.00 ± 0.14 | 0.80 ± 0.07 | 0.47 ± 0.04 | 0.000* | 0.98 ± 0.18 | 0.59 ± 0.12 | 0.53 ± 0.10 | 0.000* |
| Glucose | −0.132 | 0.201 | C2 | 1.00 ± 0.31 | 1.24 ± 0.59 | 0.31 ± 0.20 | 0.007* | 0.31 ± 0.08 | 0.15 ± 0.07 | 0.13 ± 0.06 | 0.001* |
| Xylose | −0.131 | 0.225 | C6 | 1.00 ± 0.11 | 0.85 ± 0.17 | 0.74 ± 0.06 | 0.014* | 0.76 ± 0.18 | 0.51 ± 0.12 | 0.42 ± 0.09 | 0.002* |
| Arabinose | −0.118 | 0.270 | C6 | 1.00 ± 0.15 | 1.08 ± 0.16 | 0.79 ± 0.11 | 0.012* | 0.84 ± 0.20 | 0.41 ± 0.10 | 0.42 ± 0.06 | 0.000* |
| Fructose | −0.110 | 0.236 | C2 | 1.00 ± 0.49 | 1.94 ± 0.78 | 0.44 ± 0.25 | 0.002* | 0.50 ± 0.07 | 0.19 ± 0.08 | 0.21 ± 0.07 | 0.000* |
| myo-Inositol | −0.094 | 0.226 | C2 | 1.00 ± 0.13 | 0.77 ± 0.08 | 0.71 ± 0.06 | 0.000* | 0.70 ± 0.18 | 0.71 ± 0.13 | 0.56 ± 0.08 | 0.143 |
| l-Lysine | −0.079 | 0.239 | C2 | 1.00 ± 0.13 | 1.18 ± 0.52 | 0.87 ± 0.22 | 0.349 | 1.03 ± 0.39 | 0.62 ± 0.23 | 0.61 ± 0.14 | 0.027* |
| Erythritol | −0.076 | 0.235 | C6 | 1.00 ± 0.13 | 1.07 ± 0.07 | 1.17 ± 0.14 | 0.074 | 0.17 ± 0.05 | 0.16 ± 0.06 | 0.10 ± 0.02 | 0.055 |
| Ribose | −0.031 | 0.266 | C2 | 1.00 ± 0.21 | 1.24 ± 0.30 | 1.18 ± 0.13 | 0.199 | 1.50 ± 0.35 | 0.93 ± 0.11 | 0.79 ± 0.17 | 0.000* |
| l-Tyrosine | 0.113 | −0.040 | C4 | 1.00 ± 0.51 | 1.97 ± 0.46 | 2.19 ± 0.23 | 0.001* | 1.71 ± 0.15 | 2.60 ± 0.52 | 2.69 ± 0.29 | 0.000* |
| Malate | 0.124 | 0.064 | C1 | 1.00 ± 0.23 | 2.41 ± 1.35 | 3.84 ± 0.55 | 0.000* | 0.75 ± 0.21 | 1.66 ± 0.33 | 2.23 ± 0.61 | 0.000* |
| Benzoate | 0.130 | −0.070 | C5 | 1.00 ± 0.30 | 1.71 ± 0.53 | 2.68 ± 0.95 | 0.002* | 3.76 ± 4.45 | 4.18 ± 2.05 | 3.16 ± 0.80 | 0.826 |
| 2-Oxoglutarate | 0.134 | −0.133 | C5 | 1.00 ± 0.14 | 1.28 ± 0.11 | 1.39 ± 0.19 | 0.001* | 1.47 ± 0.17 | 1.56 ± 0.27 | 1.77 ± 0.32 | 0.151 |
| Glycolate | 0.141 | 0.034 | C5 | 1.00 ± 0.20 | 1.70 ± 0.21 | 1.86 ± 0.63 | 0.004* | 1.77 ± 0.51 | 2.21 ± 0.64 | 1.71 ± 0.37 | 0.221 |
| Monomethyl-P | 0.147 | 0.130 | C3 | 1.00 ± 0.16 | 1.40 ± 0.23 | 1.69 ± 0.22 | 0.000* | 1.58 ± 0.35 | 1.68 ± 0.38 | 1.52 ± 0.34 | 0.755 |
| 4-Hydroxycinnamate | 0.148 | 0.195 | C4 | 1.00 ± 0.27 | 2.09 ± 0.72 | 2.66 ± 0.29 | 0.000* | 1.29 ± 0.47 | 1.76 ± 0.55 | 1.74 ± 0.35 | 0.174 |
| trans-Ferulate | 0.157 | 0.076 | C4 | 1.00 ± 0.55 | 2.47 ± 0.91 | 2.88 ± 0.71 | 0.002* | 2.40 ± 0.91 | 3.16 ± 0.65 | 2.75 ± 0.80 | 0.283 |
| Citrate | 0.170 | −0.073 | C4 | 1.00 ± 0.15 | 1.27 ± 0.34 | 2.21 ± 0.45 | 0.000* | 1.15 ± 0.45 | 2.28 ± 0.40 | 2.56 ± 0.21 | 0.000* |
| l-Isoleucine | 0.173 | 0.000 | C1 | 1.00 ± 0.21 | 1.47 ± 0.24 | 1.58 ± 0.14 | 0.001* | 1.41 ± 0.20 | 1.87 ± 0.36 | 1.87 ± 0.08 | 0.006* |
| trans-p-Coumarate | 0.174 | 0.161 | C4 | 1.00 ± 0.10 | 2.22 ± 0.50 | 2.65 ± 0.40 | 0.000* | 1.62 ± 0.44 | 2.04 ± 0.53 | 2.05 ± 0.25 | 0.160 |
| Erythronate | 0.176 | 0.001 | C1 | 1.00 ± 0.20 | 1.68 ± 0.20 | 2.23 ± 0.36 | 0.000* | 1.69 ± 0.38 | 2.31 ± 0.39 | 2.15 ± 0.47 | 0.050 |
| l-Glycerol-3-P | 0.176 | 0.157 | C4 | 1.00 ± 0.11 | 1.70 ± 0.27 | 2.21 ± 0.27 | 0.000* | 1.50 ± 0.30 | 1.82 ± 0.38 | 1.66 ± 0.26 | 0.235 |
| l-Serine | 0.177 | −0.019 | C1 | 1.00 ± 0.19 | 1.66 ± 0.34 | 1.82 ± 0.31 | 0.001* | 1.45 ± 0.29 | 2.45 ± 0.56 | 2.29 ± 0.17 | 0.001* |
| l-Cysteine | 0.178 | 0.014 | C1 | 1.00 ± 0.29 | 2.03 ± 0.43 | 2.50 ± 0.20 | 0.000* | 1.95 ± 0.42 | 2.62 ± 0.50 | 2.45 ± 0.53 | 0.078 |
| l-Hydroxyproline | 0.180 | 0.041 | C1 | 1.00 ± 0.23 | 1.61 ± 0.46 | 2.55 ± 0.27 | 0.000* | 1.47 ± 0.25 | 2.15 ± 0.47 | 2.15 ± 0.42 | 0.012* |
| l-Glutamine | 0.183 | −0.040 | C4 | 1.00 ± 0.30 | 4.93 ± 1.62 | 27.24 ± 4.49 | 0.000* | 1.40 ± 0.23 | 17.30 ± 5.04 | 17.03 ± 2.18 | 0.000* |
| Fumarate | 0.184 | 0.040 | C1 | 1.00 ± 0.10 | 2.37 ± 0.43 | 4.11 ± 0.24 | 0.000* | 1.42 ± 0.37 | 2.84 ± 0.74 | 3.14 ± 0.36 | 0.000* |
| l-Alanine | 0.184 | −0.011 | C1 | 1.00 ± 0.30 | 2.16 ± 0.55 | 4.40 ± 0.49 | 0.000* | 1.40 ± 0.39 | 3.75 ± 0.95 | 3.97 ± 0.53 | 0.000* |
| Threonate | 0.184 | 0.006 | C1 | 1.00 ± 0.22 | 1.96 ± 0.35 | 2.68 ± 0.32 | 0.000* | 1.89 ± 0.33 | 2.63 ± 0.45 | 2.49 ± 0.49 | 0.021* |
| l-Asparagine | 0.184 | 0.042 | C1 | 1.00 ± 0.34 | 2.55 ± 0.58 | 3.95 ± 0.70 | 0.000* | 1.42 ± 0.25 | 3.27 ± 0.91 | 3.23 ± 0.29 | 0.000* |
| l-Aspartate | 0.185 | −0.041 | C1 | 1.00 ± 0.15 | 2.38 ± 0.28 | 7.54 ± 1.17 | 0.000* | 1.27 ± 0.16 | 6.35 ± 1.60 | 5.67 ± 0.79 | 0.000* |
| Succinate | 0.186 | −0.031 | C1 | 1.00 ± 0.12 | 1.65 ± 0.30 | 2.53 ± 0.34 | 0.000* | 1.44 ± 0.16 | 2.20 ± 0.48 | 2.50 ± 0.33 | 0.000* |
| l-Homoserine | 0.186 | −0.048 | C1 | 1.00 ± 0.24 | 1.99 ± 0.64 | 3.80 ± 0.66 | 0.000* | 1.73 ± 0.62 | 4.40 ± 0.83 | 3.83 ± 0.44 | 0.000* |
| l-Glutamate | 0.187 | 0.114 | C1 | 1.00 ± 0.13 | 1.87 ± 0.24 | 2.37 ± 0.16 | 0.000* | 1.57 ± 0.21 | 2.06 ± 0.43 | 1.88 ± 0.20 | 0.038* |
| l-Threonine | 0.193 | 0.030 | C1 | 1.00 ± 0.17 | 1.83 ± 0.25 | 2.56 ± 0.38 | 0.000* | 1.48 ± 0.23 | 2.49 ± 0.51 | 2.39 ± 0.20 | 0.000* |
| Pyroglutamate | 0.193 | 0.063 | C4 | 1.00 ± 0.09 | 1.98 ± 0.25 | 3.24 ± 0.30 | 0.000* | 1.51 ± 0.31 | 2.56 ± 0.56 | 2.48 ± 0.17 | 0.000* |
| l-Valine | 0.194 | 0.058 | C1 | 1.00 ± 0.19 | 2.04 ± 0.35 | 2.85 ± 0.29 | 0.000* | 1.54 ± 0.24 | 2.66 ± 0.57 | 2.49 ± 0.37 | 0.001* |
| PC1 score | −8.70 ± 0.50 | −0.68 ± 0.64 | 4.55 ± 0.43 | −2.33 ± 0.77 | 4.12 ± 0.87 | 3.80 ± 0.36 | |||||
| PC2 score | −0.39 ± 0.59 | 2.85 ± 0.59 | 1.92 ± 0.43 | 0.40 ± 0.96 | −1.43 ± 0.98 | −3.03 ± 0.62 | |||||
| Sample number | 6 | 6 | 5 | 6 | 6 | 6 | |||||
Selected metabolites represent a high (>0.1) or low (less than −0.01) loading score of PC1. The P-value was determined by ANOVA. *P < 0.05. Compounds in bold indicate significance in both cv. Sanpia and cv. Spade one. In SOM analysis, each compound is assigned to one of the six cluster groups (C1–C6).
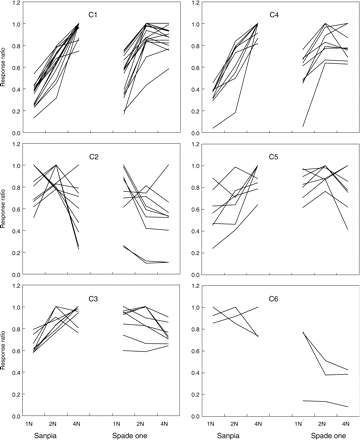
Self-organizing mapping (SOM) was used to investigate this interactive relationship for each compound (Table 2). The SOM analysis showed that among the 51 identified metabolites, 37 belonged to one of four clusters (C1, C2, C4 or C6, the membership of the cluster represented by SOM is presented in Supplement material 1). These metabolites increased or decreased, reflecting a close relationshipwith N nutrition as shown in Fig. 2. On the other hand, the remaining metabolites belonging to cluster C3 or C5 showed weaker relationships to N nutrition, at least in one cultivar (Fig. 3). In the case of 1N treatment in Sanpia, the response pattern was different from those in other treatments. The likely explanation is that because this treatment resulted in the lowest concentrations of total N and nitrate in the plant, global metabolic activity slowed down, decreasing the entire pool of metabolites.
Relative response ratio to N treatment and cultivar difference of each metabolite in spinach, cv. Sanpia and cv. Spade one, grown at different nitrogen concentrations. 1N, 2N and 4N denote 1, 2 and 4 mmol NO3 in the culture solution, respectively. The response ratio is the relative metabolite content of each compound relative to the maximum value of cultivars × N treatment.
The membership of the clusters described by SOM fitted well with the categories suggested by PCA loading data. In the case of SOM, metabolites were divided into two categories, which decreased (C2, C6) or increased (C1, C4) with N treatment (Fig. 3, Table 2). In the case of PCA, metabolites were also divided into two categories, with a positive or negative loading score (Table 2). The metabolites belonging to categories C2 or C6 by SOM correspond to the metabolites with negative loading score by PCA, while metabolites belonging to C1 or C4 by SOM correspond to those with a positive loading score by PCA (Table 2).
Discussion
To obtain a clear metabolic profile, it is important to optimize the experimental design. Data from metabolite profiling are informative enough to assess the quality of experimental design (Stitt and Fernie 2003). To begin with, successful metabolite profiling requires suitable tissues for sampling and analysis. We chose to manipulate N supply from 26 to 34 d after sowing, a period when total N accumulation is markedly increased. The most expanded leaf was chosen to obtain the required quantity of material during this limited period. The range of nitrate concentrations in the culture solution (1–4 mmol l−1) was chosen to produce a wide range of N concentrations in the tissue. Two spinach cultivars were chosen because they differed in their ability to use N. The level of total N was significantly lower in Sanpia than in Spade one (Table 1), reflecting the difference in their ability to use N, and further widening the range of tissue N concentrations in the study as a whole.
To investigate the effect of N levels on metabolites in spinach leaf, metabolite profiling was performed using GC-MS. The response of total variance of each metabolite to N concentration in the culture medium was investigated. PCA showed that the first factor accounted for 44.5% of the total variance (Fig. 1), and SOM analysis showed that 37 out of a total of 51 metabolites belonged to the categories responding to N content of the tissue (Fig. 3). Despite the plants being free of any apparent stress due to N deficiency, the N content of the tissue affected the level of most metabolites. The primary metabolites detected by GC-MS were mostly the metabolites further downstream, which are affected simultaneously by N content of the tissue. We used SOM analysis to classify the metabolites according to the pattern of their response to N. Unexpectedly, all the metabolites clustered into just two patterns: with increasing plant N, their level either increased or decreased (Fig. 3). In the case of amino acids, our data are partly supported by earlier reports showing a coordinated response of amino acids to N (Khamis et al. 1990, Noctor et al. 2002).
Comparing the N responses of the two cultivars, Sanpia exhibited lower tissue N levels than Spade one, but with a wider range: in Spade one, the tissue N level was similar between 2N and 4N treatments (Table 1). These N-dependent differences could be the primary factor explaining the different metabolite responses in the two cultivars. Several metabolites, such as p-coumarate and cysteine, showed a significant response to N treatments only in Sanpia and not in Spade one (Table 2). Thus, it appears that Sanpia shows clearer N-dependent differences with a wider range of N levels. It should be noted that those metabolites with non-significant changes in Spade one nevertheless also tended to follow a coordinated response to N. Interestingly, several metabolites showed opposite responses in the two cultivars. These compounds, such as phenylalanine (Supplement material 2), display cultivar-specific responses to N. However, our method for selecting metabolites using the PCA loading score excluded most of the metabolites without a coordinated response between two cultivars.
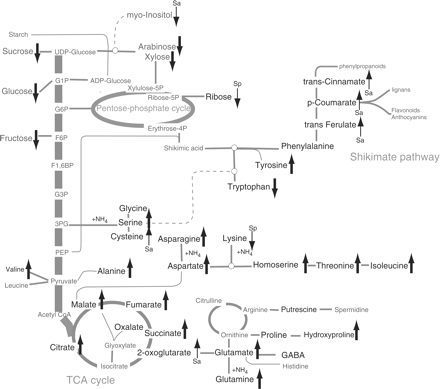
Groups of similar metabolites showed a well-coordinated response to the N status of the tissue. With the increase in N concentration in the medium, the pool of sugars decreased, whereas that of organic acids and amino acids increased. This is clearly illustrated in Fig. 4, where biosynthetic pathways have been annotated with upward or downward arrows to indicate the response to N status. It has been reported that N metabolism in plants is controlled by a complex network of hormones, nitrate, sugars, organic acids, amino acids and other chemicals (Foyer et al. 2003), and that nitrate uptake, nitrate assimilation and ammonium assimilation interact to affect metabolites further downstream (Stitt et al. 2002). Champigny and Foyer (1992) showed that nitrate redirects the flow of C away from sucrose and toward amino acid synthesis, by activating cytosolic protein kinases. Scheible et al. (1997, 2000) showed that high levels of nitrate promote organic acid synthesis by enhancing the expression of phosphoenolpyruvate carboxylase as part of the coordinated regulation of C/N. These observations are consistent with our results reported here. However, we also observed a response pattern that was specific to spinach: levels of tryptophan and lysine responded in the opposite direction to the other amino acids. A similar phenomenon has been reported in a variety of species, including beans, spinach, cauliflower and potato, whereby certain minor amino acids display an opposite response pattern to the other amino acids (Müller and Hippe 1987, Eppendorfer et al. 1996, Custic et al. 2002). The identity of the minor amino acids responding in this way varies between species; our data suggest that tryptophan and lysine are the candidates specific to spinach.
Mapping of metabolite concentrations onto plant biosynthetic pathways. Upward-pointing arrows show increased amounts under high N, and downward-pointing arrows show decreased amounts under high N. Bold arrows indicate a significant change observed in both cultivars, while narrow arrows marked Sa or Sp indicate a significant change observed only in Sanpia or Spade one, respectively. The membership of metabolites with arrows correspond to those in Table 2. Bold letters indicate identified metabolites. The metabolites which are selected in Table 2 but do not find linkage in the pathways shown in this figure are erythritol, benzoate, glycolate, monomethyl-phosphate, erythronate, threonate and pyroglutamate: these compounds are not shown.
In general, the shikimic acid pathway is promoted by N starvation (Fritz et al. 2006). However, in our experiment with spinach, most metabolites related to this pathway tended to decrease with a decrease in N (Table 2, Fig. 3). In tobacco, Fritz et al. (2006) showed that three genes, which regulate the initial steps of the shikimic acid pathway, were induced by low nitrate supply, and this may lead to increased content of chlorogenic acid, coumaric acid and caffeic acid, but not ferulic acid. The signal of N starvation represses alkaloid synthesis, which requires high N, and induces phenylpropanoid synthesis, which does not. The unique response of spinach in the present experiment was observed over a wide range of N levels and in different cultivars without any stress symptoms of N deficiency. Therefore, there may be a spinach-specific response of shikimic acid metabolism to N status, leading to changes in the levels of flavonoids, alkaloids and other phenylpropanoids.
In conclusion, this study showed that different levels of N led to differences in metabolite profiles. With changing tissue N content, a marked change was observed in the metabolite composition of carbohydrates, organic acids and amino acids, which together represented 44.5% of the total variance in PCA analysis. Both global and spinach-specific changes in metabolite composition were detected by GC-MS. These results indicate the utility and importance of metabolite profiling by GC-MS in studies investigating the composition of primary metabolites such as carbohydrates, organic acids and amino acids. Based on our results, we suggest that greater attention should be paid to the overall relationship between plant N status and metabolites when monitoring environmental effects on plants or investigating characteristics that relate to C/N status.
Materials and Methods
Plant growth
Seeds of spinach (S. oleracea L., cv. Sanpia and cv. Spade one) were germinated, and the seedlings were grown hydroponically in 3 liter pots under a light intensity of 630 μmol photons m−2 s−1 at 23/18°C under a 11/13 h light/dark regime, in a growth room at the National Agricultural Research Center for Hokkaido Region. The composition of the nutrient solution was as follows: 0.4 g l−1 KNO3, 0.09 g l−1 NaH2PO4·2H2O, 0.15 g l−1 Na2HPO4·12H2O, 0.29 g l−1 CaCl2·2H2O, 0.49 g l−1 MgSO4·7H2O, 23 mg l−1 Fe-EDTA, 2.9 mg l−1 H3BO3, 1.8 mg l−1 MnCl2·4H2O4, 0.22 mg l−1 ZnSO4·7H2O, 0.08 mg l−1 CuSO4·5H2O and 0.03 mg l−1 Na2MoO4·2H2O. Three N treatments (1, 2, and 4 mmol l−1 NO3-N, denoted 1N, 2N and 4N, respectively) were applied about 8 d before harvest. The most expanded leaf was harvested from each plant 34 d after sowing, 1–1.5 h after the beginning of the light period. The leaves were flash-frozen under liquid nitrogen, lyophilized, and stored at −80°C until needed.
Determination of nitrogen
Total N and nitrate concentrations were determined in whole leaves of spinach. Nitrate was analyzed by ion-chromatography (Dionex, ICS-90). Total N was analyzed by the auto analyzer method (Bran + Luebbe, AACS-III), after Kjeldahl decomposition.
Extraction and derivatization
Metabolite analysis was carried out according to the method of Roessner et al. (2000). Briefly, leaf tissue (10 mg) was mashed in a 2 ml tube using a multibead shocker (Yasuikikai, Osaka, Japan) at 2000 (smash intensity) for 10 s (Sato et al. 2004). Ice-cooled methanol (300 μl) and subsequently ribitol (30 μl, 0.02 mg ml−1 water) was added to the mashed tissue. The sample was mixed and incubated for 15 min at 70°C. Chloroform (200 μl) was added to the solution and incubated for 5 min at 37°C. Next, 400 μl of water was added to separate the polar and non-polar phases. After centrifugation (20,000 × g, 5 min, 4°C), 10 μl of upper methanol/water phase was transferred to each autosampler vial and dried in a vacuum centrifuge at 15°C.
Samples were automatically derivatized by 10 μl of methoxyamine hydrochloride (90 min, 40°C) and 17.5 μl of N-methyl-N-trifluoroacetamide (MSTFA; 30 min, 40°C) with Combi-PAL (CTC Analytics), according to the method described by Erban et al. (2007). A 2.5 μl aliquot of a retention time standard mixture [n-decane, n-dodecane, n-pentadecane, n-octadecane, n-nonadecane, n-docosane, n-octacosane, n-dotriacontane and n-hexatriacontane (0.029%, v/v, in pyridine)] was added with MSTFA. A 1 μl aliquot of the sample was injected into a gas chromatograph (Agilent GC 6890) in the splitless mode. Gas chromatography was performed on an Rtx-5Sil MS with an integrated guard column (30 m, 0.25 μm film; Restek GmbH, Bad Homburg, Germany). The injection, interface and ion source temperatures were adjusted at 230, 250 and 210°C, respectively. The gas flow rate was 1 ml min−1. The column temperature was held for 1 min at 70°C, 6 min up to 76°C, 45 min up to 350°C, 1 min at 350°C and 10 min at 330°C. The column end was introduced into a GCmate-II sector mass spectrometer (JEOL, Tokyo, Japan). The mass spectra were recorded at 2 scan s−1 with a m/z 50–600 scanning range. Metabolites were identified by mass spectral and retention index using AMDIS software (http://chemdata.nist.gov/mass-spc/amdis/), referencing a private library of 183 self-purchased standards including all annotated metabolites in the Golm metabolome database (http://csbdb.mpimp-golm.mpg.de/csbdb/gmd/gmd.html), and the metabolites belonging to the pathways shown in Fig. 4. Identified metabolites were quantified using Quant software (JEOL, Tokyo, Japan), following the process described by Roessner et al. (2001). Before statistical analysis, the data were normalized using the peak area of ribitol.
Standardization of each analysis
A SOM analysis was performed using Viscovery SOMine (Eudaptics software Gmbh, Austria). After log10 data transformation, PCA was carried out using SIMCA-P 11.0 (Umetrics AB). The matrix of 51 metabolites × 36 biological individuals was used for both analyses.
Supplementary material
Supplementary material mentioned in the article is available to online subscribers at the journal website www.pcp.oxfordjournals.org.
Funding
The Ministry of Education, Science, Sports and Culture's Grant-in-Aid for Scientific Research (B) (17658144, 2005).
Acknowledgments
We would like to thank Dr. Oliver Fiehn (UC Davis Genome Center) for useful suggestions regarding GC-MS analysis, Dr. Miyako Kusano and Makoto Kobayashi (RIKEN Plant Science Center) for providing technical details, and Dr. Alexander Erban and Dr. Joachim Kopka (Max Planck Institute) for providing details of the methods for automated chemical derivatization with Combi-PAL. We thank Dr. T. Nakamura (NARO) for his helpful comments on and discussion of our recent studies. We also thank Mr. Naoki Matsumoto and Ms. Iu Sunaga for technical assistance in operating GC-MS.
References
Abbreviations:
- GC-MS
gas chromatography–mass spectrometry
- MSTFA
N-methyl-N-trifluoroacetamide
- PCA
principal component analysis
- SOM
self-organizing mapping.